 The way that transcription factors of the Hox class, particularly those encoded by the homeotic genes, are able to direct the development of specific animal morphologies is a long-standing question in developmental biology. Many years of developmental genetics and molecular biology, starting from the Nobel Prize winning work of Ed Lewis, established much about how Hox factors control development, however, we still have a relatively poor understanding of how individual Hox proteins operate at a genome wide scale.
The way that transcription factors of the Hox class, particularly those encoded by the homeotic genes, are able to direct the development of specific animal morphologies is a long-standing question in developmental biology. Many years of developmental genetics and molecular biology, starting from the Nobel Prize winning work of Ed Lewis, established much about how Hox factors control development, however, we still have a relatively poor understanding of how individual Hox proteins operate at a genome wide scale.
This is particularly pertinent since all of the homeotic factors characterised to date in the Fly recognise very similar DNA sequences (TAAT) thus the question of how the specificity that allows elaboration of different structures along the body axis remains somewhat enigmatic. We have begun to take a genomics approach to this question, building on our long-stabnding collaboration with Rob White’s lab. With the serendipitous recovery of protein traps in both Ultrabithorax (Ubx) and its known cofactor Homothorax (Hth) in our protein trap screen, we were able to carry out a genome wide analysis of Ubx and Hth binding in the haltere, wing (Hth) and leg imaginal discs – work by Lawrence Choo published in PLoS ONE . A similar analysis form Richard Mann and Kevin White (Slattery et al) generated similar data, leading us to conclude we have a reasonably accurate picture of genome wide Ubx binding in a single tissue.
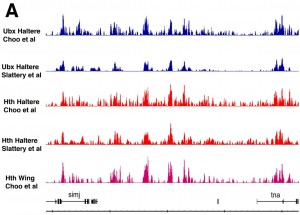 Our interpretation of these data are that aspects of chromatin accessibility may be a large driving force behind Hox protein binding in the genome and raise questions about how specificity is achieved. The idea that chromatin accessibility direct TF binding in the genome is supported by the considerable genome wide analysis of TF binding and chromatin features in the early Drosophila embryo by the Berkley Drosophila Transcriptional Network Project and may prove to be a general feature of gene regulation in metazoans.
Our interpretation of these data are that aspects of chromatin accessibility may be a large driving force behind Hox protein binding in the genome and raise questions about how specificity is achieved. The idea that chromatin accessibility direct TF binding in the genome is supported by the considerable genome wide analysis of TF binding and chromatin features in the early Drosophila embryo by the Berkley Drosophila Transcriptional Network Project and may prove to be a general feature of gene regulation in metazoans.
We are currently investigating the effects of ectopic Hox expression at a genome wide scale with a view to understanding how specificity is achieved. We are doing this both in the animal and in a cell culture model.
Current Lab Members
Former Lab Members
Collaborators