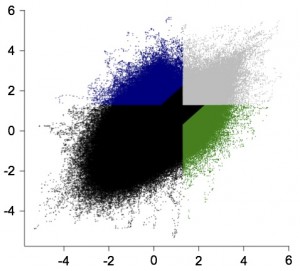
 A landmark for the lab with our first ever BioConductor package now available. SimBindProfiles allows comparisons of ChIP-array binding profiles to identify differential binding in different datasets, while there are tools to this with ChIP-seq data there was little or nothing usable for array based analysis. Bettina put it together with some help from Enrico and Robert Stojnic to facilitate our work on Dichaete and SoxN binding but it is broadly applicable to any ChIP-array datasets.
A landmark for the lab with our first ever BioConductor package now available. SimBindProfiles allows comparisons of ChIP-array binding profiles to identify differential binding in different datasets, while there are tools to this with ChIP-seq data there was little or nothing usable for array based analysis. Bettina put it together with some help from Enrico and Robert Stojnic to facilitate our work on Dichaete and SoxN binding but it is broadly applicable to any ChIP-array datasets.
I think this is a fabulous piece of work – well done Bettina!
Leave a Reply